Regulation of the Mammalian Elongation Cycle by Subunit Rolling: A Eukaryotic-Specific Ribosome...
-
Upload
charite-de -
Category
Documents
-
view
2 -
download
0
Transcript of Regulation of the Mammalian Elongation Cycle by Subunit Rolling: A Eukaryotic-Specific Ribosome...
Regulation of the Mammalian ElongationCycle by Subunit Rolling: A Eukaryotic-Specific Ribosome RearrangementTatyana V. Budkevich,1,2,3 Jan Giesebrecht,1 Elmar Behrmann,1 Justus Loerke,1 David J.F. Ramrath,1 Thorsten Mielke,1,4
Jochen Ismer,1 Peter W. Hildebrand,1 Chang-Shung Tung,5 Knud H. Nierhaus,1,2 Karissa Y. Sanbonmatsu,5,6
and Christian M.T. Spahn1,*1Institut fur Medizinische Physik und Biophysik, Charite–Universitatsmedizin Berlin, Chariteplatz 1, 10117 Berlin, Germany2Max-Planck Institut fur Molekulare Genetik, Abteilung Vingron, AG Ribosomen, 14195 Berlin, Ihnestraße 73, Germany3Institute of Molecular Biology and Genetics, Group of Protein Biosynthesis, 03143 Kiev, Ukraine4Max-Planck Institut fur Molekulare Genetik, UltraStrukturNetzwerk, 14195 Berlin, Ihnestraße 73, Germany5Theoretical Biology and Biophysics Group, Theoretical Division, Los Alamos National Laboratory, MK710, Los Alamos, NM 87545, USA6New Mexico Consortium, 4200 West Jemez Road, Suite 301, Los Alamos, New Mexico 87544, USA
*Correspondence: [email protected]
http://dx.doi.org/10.1016/j.cell.2014.04.044
SUMMARY
The extent to which bacterial ribosomes and thesignificantly larger eukaryotic ribosomes sharethe same mechanisms of ribosomal elongation isunknown. Here, we present subnanometer resolut-ion cryoelectron microscopy maps of the mam-malian 80S ribosome in the posttranslocational stateand in complex with the eukaryotic eEF1A,Val-tRNA,GMPPNP ternary complex, revealing sig-nificant differences in the elongation mechanismbetween bacteria and mammals. Surprisingly, andin contrast to bacterial ribosomes, a rotation of thesmall subunit around its long axis and orthogonalto the well-known intersubunit rotation distinguishesthe posttranslocational state from the classical pre-translocational state ribosome. We term this motion‘‘subunit rolling.’’ Correspondingly, a mammaliandecoding complex visualized in substates beforeand after codon recognition reveals structural dis-tinctions from the bacterial system. These findingssuggest how codon recognition leads to GTPaseactivation in the mammalian system and demon-strate that in mammalia subunit rolling occurs duringtRNA selection.
INTRODUCTION
Genetic informationwithinmRNA is translated into protein during
the elongation phase of translation (Voorhees and Ramak-
rishnan, 2013). Ribosomes decode one codon of the mRNA
sequence per elongation cycle by using tRNA substrates and
append the encoded amino acid to the nascent peptide. An elon-
gation cycle can be subdivided into three steps: (1) delivery of
aminoacyl-tRNA (aa-tRNA), which involves decoding and ac-
commodation, (2) peptide-bond formation, and (3) tRNA translo-
cation. Peptide-bond formation is catalyzed by the ribosome’s
peptidyl transferase center and is a fast and spontaneous step.
However, during decoding and translocation the ribosome has
to overcome large activation energy barriers. Accordingly, the
pretranslocational (PRE) and the posttranslocational (POST)
states of the ribosome are relatively stable (Schilling-Bartetzko
et al., 1992). These activation energy barriers are reduced by
translational GTPase elongation factors, which are responsible
for speed and accuracy of protein synthesis.
The bacterial elongation cycle and its substeps have been
extensively studied over the past decades using a battery of
functional, genetic, and structural methods (Frank and Spahn,
2006; Voorhees and Ramakrishnan, 2013). An elongation cycle
starts when an aa-tRNA is delivered to the POST state ribosome
carrying a peptidyl-tRNA in the P-site and a deacylated tRNA
in the E-site. A-site occupation is a complex, multistep proc-
ess (Schmeing et al., 2009; Schuette et al., 2009; Voorhees
and Ramakrishnan, 2013). In the initial decoding step the aa-
tRNA,EF-Tu,GTP ternary complex binds to the ribosome.
Recognition of the cognate codon leads to a complex rearrange-
ment of the tRNA. In the resulting A/T-state the tRNA can bind to
the A-site of the ribosomal 30S subunit and remains simulta-
neously bound to EF-Tu, which in turn interacts with the ribo-
some’s factor-binding site. The conformational rearrangements
of the ternary complex, combined with changes in the ribosome,
transmit decoding signals from cognate codon-anticodon inter-
actions to EF-Tu-stimulating GTP hydrolysis and subsequent
dissociation of EF-Tu,GDP. Decoding is completed when the
body of the tRNA swings into the ribosome through the accom-
modation corridor and establishes interactions with the A-site of
the 50S subunit during accommodation (Sanbonmatsu et al.,
2005). After peptide-bond formation, the resulting PRE complex
contains a peptidyl-tRNA in the A-site and a deacylated tRNA in
the P-site. In order to reset the ribosome for the next round of
elongation, the EF-G-dependent translocation reaction moves
Cell 158, 121–131, July 3, 2014 ª2014 Elsevier Inc. 121
the tRNA2,mRNA complex by one codon through the ribosome,
establishing the POST state (Voorhees and Ramakrishnan,
2013).
While the structure and function of the ribosome is sub-
stantially conserved across the domains of life, comparatively
little is known about the detailed mechanism of eukaryotic
translation. The eukaryotic (80S) ribosome is significantly lar-
ger and more complex than its bacterial counterpart (Anger
et al., 2013; Ben-Shem et al., 2011; Klinge et al., 2011; Spahn
et al., 2004a) and mechanistic investigations of the transla-
tion mechanisms in higher organisms are lacking. Substantial
disparities in the translation mechanism between eukaryotic
and prokaryotic systems are implied by the elaborated initia-
tion, termination, and recycling mechanisms, requiring num-
erous additional factors in eukaryotes (Melnikov et al., 2012).
In contrast, the elongation phase is thought to be highly
conserved across domains of life because core elements of
the ribosome, including the substrate-binding sites and the
general elongation factors, are largely conserved. Conversely,
the existence of ribosome-targeting antibiotics that display
domain specificity (Wilson, 2009) and evidence of essential,
domain-specific translation factors (Andersen et al., 2006), indi-
cates that aspects of the translation mechanism must differ. A
first analysis of the mammalian PRE complex revealed distinc-
tions of the mammalian 80S ribosome with respect to the dy-
namic behavior of the complex and the exact nature of the
tRNA-binding sites (Budkevich et al., 2011). Here, we explore
the mammalian elongation cycle through structural investiga-
tions of the mammalian 80S ribosome decoding complexes
and the POST state from rabbit liver using cryoelectron micro-
scopy (cryo-EM). Strikingly, substantial differences to analo-
gous bacterial complexes can be observed, revealing that the
bacterial and the mammalian elongation cycle diverge more
than previously thought.
RESULTS
Cryo-EM Maps for Mammalian POST, PRE,and Decoding ComplexesIn order to provide the structural foundation for the A-site
occupation/decoding step in the mammalian system, we
used cryo-EM to analyze ribosomal decoding and POST com-
plexes. Both specimens were prepared in vitro from mamma-
lian components (Budkevich et al., 2011; Budkevich et al.,
2008). To yield the decoding complex the ternary complex
Val-tRNA,eEF1A,GMPPNP was stalled by the nonhydrolyzable
GTP analog. For the POST complex, PRE ribosomes were
translocated by eEF2 to move N-acylated Lys-tRNALys3 and
deacylated tRNAPhe from A- and P-sites to P- and E-sites,
respectively (Experimental Procedures).
The resulting complexes were analyzed by multiparticle cryo-
EM (Loerke et al., 2010). The compositional heterogeneity of the
POST specimen was larger than expected from the biochemical
data and the subpopulation corresponding to the desired 80S
POST complex consisted of only about 35% of ribosomal com-
plexes (Figure S1A available online and Extended Experimental
Procedures). This major subpopulation (236,113 particle images)
representing a POST complex with two tRNAs in classical P- and
122 Cell 158, 121–131, July 3, 2014 ª2014 Elsevier Inc.
E-sites (Figure 1A) was further refined to a resolution of 6.9 A
(Figure S1B).
The classical PRE state is the final product of the ribosomal
A-site occupation/decoding step. We made use of the presence
of significantly populated classical PRE states within the POST
data set (Figure S1A) to furthermore obtain cryo-EM maps of
the 80S ribosome in classical-1 (66,618 particle images) and
classical-2 (73,951 particle images) PRE states (Figures 1B and
1C) at 7.6 A and 7.5 A resolution, respectively (Figure S1B).
Both maps exhibited density for tRNAs in classical A-, P-, and
E-sites. They agree well with our previously reported structures
(Budkevich et al., 2011) but are significantly improved in terms
of map quality and resolution. Moreover, the PRE and the
POST maps were obtained from the same sample, thus facili-
tating a direct comparison of both states. For the ribosomal de-
coding complex, our multiparticle approach resulted in two
maps (Figures 1D and 1E) in subtly different conformations
(see below) with resolutions of 8.7 A and 8.9 A, respectively
(Figure S1B).
In total, we obtained five cryo-EM maps delineating the de-
coding process from the POST state via two decoding states
to the classical-1 and -2 PRE states (Figure 1). All maps show
typical features expected at subnanometer resolution such as
major and minor grooves of RNA double helices, a helices, and
extended protein tails. To interpret our maps in molecular terms,
we created a cryo-EM-based homology model of the mamma-
lian 80S ribosome. As secondary structure maps for rRNAs
from rabbit are not available, and rabbit and human ribosomes
are very similar (Budkevich et al., 2011; Spahn et al., 2004b),
we chose the human ribosome as target for our modeling. The
high-sequence conservation (see Extended Experimental Pro-
cedure) also suggests that a human-based model is a valid
approximation for the presented cryo-EM maps. Importantly,
our model will aid subsequent studies in the human system.
Recent X-ray structures of the ribosomal 40S and 60S subunits
from Tetrahymena thermophila (Klinge et al., 2011; Rabl et al.,
2011) and the yeast 80S ribosome (Ben-Shem et al., 2011) allow
homologymodeling not only for the evolutionary conserved inner
core of the ribosome but also for many of the eukaryotic-specific
components (Figure 1A, Table S1). Recently, a model for the
human 80S ribosome in the rotated state became available
based on a cryo-EM map at 5.4 A resolution (Anger et al.,
2013). The model presented here is in good overall agreement
with this model regardless of our use of an alternative sequence
for the ribosomal RNAs.
The Path of the mRNA in Eukaryotic Ribosomes andCodon-Anticodon Interactions with P- and E-Site tRNAsThe presented cryo-EM map of the mammalian POST complex
allows a direct visualization of an mRNA fragment with a length
of approximately 34 nucleotides (Figures 1A, 2A, and 2B). The
modeled mRNA ranges from position -15 on the 50 side to posi-
tion +19 on the 30 side. This agrees well with the part of the
mRNA that is shielded by the ribosome according to mRNA pro-
tection studies (Steitz, 1969) and recent ribosome profiling data
(Ingolia et al., 2011). As in bacterial ribosomes (Jenner et al.,
2010), the mRNA enters the mRNA groove of the 40S subunit
from the solvent side through the mRNA entry tunnel, wraps
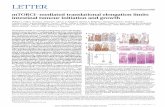
Figure 1. Reconstruction of Eukaryotic
80S POST, Classical-1 PRE, Classical-2
PRE, Codon Sampling and Codon Recogni-
tion/GTPase Activation Complexes from
Rabbit Liver
(A–E) Reconstruction of eukaryotic 80S POST (A),
PRE (classical-1) (B), PRE (classical-2) (C), codon
sampling (D), and codon recognition/GTPase
activation (E) complexes from rabbit liver. Left:
Overall view of the cryo-EM reconstructions, dis-
playing the 60S subunit (blue), 40S subunit (yel-
low), A-tRNA (pink), P-tRNA (green), E-site tRNA
(orange), A/T-tRNA (dark violet), eEF1A (red), and
mRNA (blue). Middle and right: Individual mesh
representation of the subunit maps with docked
models, showing the ribosomal ligands relative to
the 40S subunit (ribosomal RNA yellow and S
proteins gray) and the 60S subunit (rRNA blue and
L proteins orange), respectively. Landmarks are
denoted for 40S: beak (bk), left foot (lf), right foot
(rf), head (h), shoulder (sh), and 60S: central pro-
tuberance (CP), L1 stalk (L1), stalk base (SB) and
stalk (St). See also Figure S1 and Table S1.
around the neck of the 40S subunit, where it interacts with the
tRNAs, and leaves the 40S through the mRNA exit tunnel. On
the 50 side the overall path of the mRNA through the mRNA
exit tunnel deviates significantly from the bacterial system (Jen-
ner et al., 2010), whereas on the 30 side, the path of the mRNA
through the mRNA entry tunnel is remarkably similar (Figure 2B).
However, there is a kink in the mRNA at the solvent side of the
entry tunnel and about four nucleotides of mRNA (+16 to +19),
which are not visible in the bacterial system, lead upward toward
the 40S head and 18S rRNA helix 16 (h16).
In the P-site the three-nucleotide helix formed by codon-anti-
codon interaction can be directly observed (Figure 2A). Interest-
ingly, we also observe a shorter interaction interface between the
mRNA codon and the tRNA anticodon in the E-site, indicating
Cell 158, 121
one or at most two base pairing inter-
actions (Figure 2A). This is consistent
with bacterial X-ray structures where
Watson-Crick base pairing of the E-site
tRNA anticodon with the first nucleotide
of the E-site codon was described (Jen-
ner et al., 2010). Thus, codon-anticodon
interaction at the E-site may be a feature
of the mammalian POST complex, but
weakened with respect to the P-site.
Interactions of tRNA and mRNAwith Ribosomal Proteins Specificto EukaryotesThe domain-specific differences in the
path through the mRNA exit tunnel on
the mRNA’s 50 side can be explained by
the presence of the eukaryotic-specific
ribosomal proteins eS26 and eS28 (we
use the new system for naming ribosomal
proteins according to Ban et al., 2014).
The N- and C-terminal parts of eS26 block the bacterial path of
the mRNA and at the same time shield the 30-end of 18S rRNA
preventing formation of Shine-Dalgarno-like interactions. More-
over, eS26, eS28, and also uS11 (rpS14) appear to interact with
the eukaryotic mRNA and thus line out an alternative path of the
mRNA exit (Figure 2B).
At the solvent side of themRNA entry tunnel, a eukaryotic-spe-
cific contact with the 30 side of the mRNA may also take place,
facilitated by the C-terminal part of protein eS30. Interestingly,
eS30 wraps around the shoulder, and reaches into the decoding
center (Rabl et al., 2011), where its N terminus has been pro-
posed to interact with the A-site tRNA in the mammalian PRE
complex (Budkevich et al., 2011). Thus, eS30 could provide a
structural link between the outer and inner ends of the mRNA
–131, July 3, 2014 ª2014 Elsevier Inc. 123
Figure 3. 40S Subunit Rolling
(A and B) Comparison of the 40S subunit positions in classical-1 state (orange)
to the POST state (yellow) represented by (A) cryo-EM maps and (B) ribbons.
Comparisons are based on a common 60S alignment. Arrows indicate the
direction of movement during transition between the two different states. The
distance changes in the 40S subunit positions resulting from the rigid body
transformation are color-coded in A units. Landmarks for 40S are denoted:
head (h), body (b) and beak (bk). See also Figure S3 and Movie S1.
Figure 2. Eukaryotic-Specific Features of the mRNA Path Visible
from Intersubunit Space and Solvent Side of 40S Subunit
(A) Interaction of the P- and E-site tRNAs of the 80S POST complex (trans-
parent gray) with mRNA.
(B) Comparison of eukaryotic (transparent gray) and prokaryotic (red ribbon,
Jenner et al., 2010, PDB ID 3I8G) mRNA paths. The model for the additional
four nucleotides on the 30-end of the eukaryotic mRNA is shown in cyan.
Densities for the ligands were segmented from the cryo-EM map presented in
Figure 1A. The models for tRNAs, mRNA and ribosomal proteins from the
homology model of the human 80S ribosome are presented in this paper.
Ribosome orientations are indicated by orientation aids.
See also Figure S2.
entry tunnel. Hence, our structure suggests a functional role for
the eukaryotic ribosomal proteins uS11, eS26, eS28, and eS30
in escorting the mRNA through the 40S subunit. Furthermore,
eukaryotic-specific contacts also contribute to the tRNA-binding
sites (Figure S2).
40S Subunit Rolling: A Novel Mode of IntersubunitRearrangementThe presented POST complex contains classically configured
tRNAs in the P/P- and E/E-sites (Figure 1A). It is therefore ex-
pected to correspond, in terms of overall ribosome conforma-
tion, to the classical-1 PRE state that also carries classically
configured tRNAs (Figure 1B). Unexpectedly, the subunit
arrangement of the POST state differs markedly from that of
the classical-1 PRE state (Figure 3A, Movie S1). The underlying
conformational change, which we term ‘‘subunit rolling,’’ can
be described as a �6� rotation of the 40S subunit toward the
L1 stalk around the long axis of the small subunit. The axis co-
localizes approximately with the upper part of h44 of 18S rRNA
and is roughly orthogonal to the well-known intersubunit rotation
(Figure S3A).
Interestingly, subunit rolling shapes the openings to the inter-
subunit space and causes reciprocal opening and closing of
the A- and E-site regions (Movie S1). The distance between
the 40S and 60S subunits on the A-site side of the 80S rib-
osome decreases by about 13–15 A during subunit rolling
from the POST state to the classical-1 PRE state. As a conse-
quence, the A-site region is more widely open in the POST
state. The opposite is observed for the E-site, which is narro-
wer in the POST state than in the classical-1 PRE state. How-
ever, due to the smaller distance from the rotation axis, the
underlying movements are only in the range of 6–7 A at the
E-site. Subunit rolling also affects the interactions between
124 Cell 158, 121–131, July 3, 2014 ª2014 Elsevier Inc.
the 40S and the 60S subunit, i.e., the intersubunit bridges
(Figure S3D). Most of the conserved bridges are present in
the current POST 80S state with the exception of B6 and B7
(Figure S3D, in green), which are, however, found in the
mammalian classical-1 PRE state. Subunit rolling provides an
explanation for this observation as it results in movement on
the order of �5–7 A in the lower part of 40S (Figure 3B). More-
over, subunit rolling is expected to affect tRNA positions.
Indeed, the elbow region of the P/P-tRNA in the PRE state is
shifted by �6 A toward the E-site in comparison with the
mammalian POST state (and also the bacterial PRE state; Fig-
ures S3E and S3F). Thus, some differences in tRNA positioning
exist for the mammalian 80S ribosome between the classical
PRE and POST states, with either a deacylated or peptidyl-
tRNA being present in the P-site, respectively.
Cryo-EM Analysis of the Mammalian Decoding ComplexThe observed difference in the subunit arrangement of the 80S
ribosome in POST and classical PRE states has consequences
for themechanismof tRNAselection. It implies that subunit rolling
has to occur when the POST complex is converted into the PRE
complex. To exploremammalian A-site occupation, we analyzed
a mammalian decoding complex. Unexpectedly, we were able
to observe two subpopulations of the 80S,Val-tRNA,eEF1A,GMPPNP complex (Figures 1D and 1E). Both cryo-EM maps
display clear density for the ternary complex (Figures 4A and
4B) but are distinguished by more subtle changes in position,
conformation and interaction patterns between the ternary com-
plex and the 80S ribosome (Figures 4, 5, 6, and S4 and S5).
In terms of subunit configuration, both substates of the
80S,Val-tRNA,eEF1A,GMPPNP complex are similar to the
POST complex. When the larger substate of the decoding com-
plex (79,705 particle images) and the smaller one (52,686 particle
images) are compared to the POST complex only small rotations
of �0.5� and �1�, respectively, between the 40S subunits were
Figure 4. Conformation of the Mammalian
Ternary Complex and Differences from the
Bacterial Counterpart
(A and B) Overall fitting of crystallographic mo-
dels of Aeropyrum pernix aEF1A (Kobayashi
et al., 2012) (red ribbon, PDB ID 3VMF)
and T. thermophilus A/T-tRNA (pink ribbon,
modified from PDB IDs 2XQD and 1TTT for ASL
and tRNA body, respectively) to the ternary com-
plex cryo-EMmaps (transparent gray) of (A) codon
sampling state and (B) codon recognition/GTPase
activation state. Densities for the ternary complex
are extracted from the cryo-EMmaps presented in
Figure 1D and E.
(C–E) Superposition of the codon sampling state
(transparent gray) and codon recognition /
GTPase activation state ternary complex molec-
ular models on the 40S subunit surface (C). The
alignment was based on the 40S subunit densities.
(D and E) Comparisons of the mammalian
(D) codon sampling state and (E) codon recogni-
tion/GTPase recognition state (E) ternary com-
plexes with the bacterial ternary complex stalled
by GTP analog (Voorhees et al., 2010) (PDB ID
2XQD; transparent gray). The alignment was
based on conserved parts of the 18S/16S rRNA.
Ribosome orientations are indicated by orientation
aids. (C–E) The distances between positions of
the ternary complexes are color coded (capped at
6 A). We note that these distances essentially
reflect rigid body transformations of eEF1A/EF-Tu
and ASL or body of tRNA.
See also Figure S4 and Movie S2.
found. In contrast, comparing the two subpopulations of the
decoding complex with the classical-1 PRE complex reveals
respective rotations of �5.6� and �5.0�. Thus, subunit rollingoccurs after the decoding step during accommodation of the
tRNA from the A/T- to the A/A-state. By contrast, structural in-
vestigations in bacteria did not show evidence for such a major
subunit rearrangement (Schmeing et al., 2009; Schuette et al.,
2009; Voorhees et al., 2010).
Conformational Heterogeneity of the MammalianDecoding ComplexFor a molecular interpretation of the mammalian decoding com-
plexes we used the X-ray structure of aEF1A from the archaeal
EF1A/RF1 complex (Kobayashi et al., 2012), which we could fit
well as a rigid body into the cryo-EM maps of both subpopula-
tions (Figures 4A and 4B). Archaeal aEF1A and human eEF1A
Cell 158, 121
are closely related by sequence (Fig-
ure S4A). Remarkably, density for the
only major insertion helix 5 (aa 214–225)
(human numbering), which is present in
eEF1A but not in aEF1A, is observed in
our cryo-EM map and creates a signifi-
cant mismatch between model and map
at the C-terminal part of the G domain
of eEF1A before the junction to domain II
(Figure S4B). To account for the bend
between the anticodon-stem loop (ASL) and the D-stem, the
structure of the tRNA was flexibly docked (Figures 4A and 4B).
A comparison between both subpopulations of the mamma-
lian 80S,Val-tRNA,eEF1A,GMPPNP complex reveals a rota-
tional movement of eEF1A (and tRNA) around a hinge at the
interaction between the two evolutionary conserved 258GIGTV
and 289VKS/312VKN loops (human numbering) of eEF1A domain
II with the shoulder region of the 40S subunit (Figure 5A and
Movie S2). It closes an apparent gap between the G domain
(domain I) of eEF1A and the sarcin-ricin loop (SRL, H95 of 28S
rRNA) so that interactions between both elements can be
observed only in the smaller subpopulation (Figure 5B, right).
With 12 nucleotides, the SRL is the longest universally conserved
sequence of rRNAs and an essential part of the factor-binding
site. Because interactions between the SRL and the switch re-
gions of EF-Tu have been assigned a prominent role in the
–131, July 3, 2014 ª2014 Elsevier Inc. 125
Figure 5. Interactions of the Ternary Com-
plexes with the 80S Ribosome
(A) Contacts of eEF1A with the shoulder of the 40S
ribosomal subunit.
(B and C) Contacts of (B) eEF1A and (C) A/T-tRNA
inside of the ternary complexes with SRL of 28S
rRNA.
(D) Contact of the eEF1A domain III with the C
terminus of uS12. Left column represents the
codon sampling state, right column represents
the codon recognition / GTPase activation state.
Ribosome orientations are indicated by orienta-
tion aids. The UniProt numbering which includes
the leading methionine, was used, resulting in
a +1sequence shift, such that e.g., eukaryotic
His94 is His95 according to our numbering.
See also Figure S5.
molecular mechanism of GTPase activation in the bacterial sys-
tem (Voorhees and Ramakrishnan, 2013), we propose that the
smaller subpopulation of the 80S,Val-tRNA,eEF1A,GMPPNP
complex is at least close to the GTPase activation state.
This interpretation is corroborated by the appearance of the
ribosomal decoding center in our cryo-EM maps (Figure 6). For
the smaller subpopulation of the decoding complex (Figure 6D)
as well as for the classical-2 and -1 PRE complexes (Figures 6E
and 6F, respectively) the respective cryo-EM densities are
compatible with a fully flipped-out conformation of the bases
A1824 and A1825 of 18S rRNA at the top of h44 (A1492/A1493
in E. coli) allowing minor groove interactions with the codon-anti-
codon helix of the A/T- or A-tRNA. These minor-groove interac-
tions are at the heart of the ribosomal decoding mechanism and
126 Cell 158, 121–131, July 3, 2014 ª2014 Elsevier Inc.
ensurefidelity of tRNAselection (Voorhees
and Ramakrishnan, 2013) (Figure 6A).
Because codon-recognition is a prerequi-
site for GTPase activation, the SRL inter-
actions and the state of the decoding
center both suggest that the smaller sub-
population of our mammalian 80S,Val-tRNA,eEF1A,GMPPNP complex repre-
sents the codon-recognition/GTPase
activation state of the mammalian sys-
tem, and is functionally equivalent to
the bacterial structure of the GDPCP
stalled 70S,tRNA,EF-Tu complex (Voo-
rhees et al., 2010).
In contrast, the POST complex (Fig-
ure 6B) and the larger substate of the
decoding complex (Figure 6C) do not
display density for stably flipped-out
18S rRNA bases A1824/A1825 indicating
a more dynamic state of the top of h44.
Therefore, and in line with the observa-
tion that eEF1A does not yet interact
with the SRL (Figure 5B, left), we suggest
that the larger substate of themammalian
decoding complex represents an initial
codon sampling state. Here, codon-anti-
codon interaction occurs but is not yet stabilized. So far such a
state of the ribosomal decoding complex has not been directly
visualized.
Concomitant with the occupation of the 40S decoding center
by an ASL and the stabilization of the codon-anticodon duplex
by the flipped-out 18S rRNA bases A1824/A1825, a progressive
tightening of the interaction surface between the top of 18S rRNA
h44 and the shoulder region (h18 and uS12) can be observed
(Figure 6A). These more local changes appear embedded in an
overall tightening of the 40S subunit around the ASL, which in-
volves a subtle movement of the shoulder region, in particular
h16, and the 40S head (Movie S3). These changes are reminis-
cent of the bacterial domain closure movement of the 30S sub-
unit (Ogle et al., 2002). However, the mammalian conformational
Figure 6. Codon Recognition and 40S
Domain Closure at the Decoding Center
(A) Models for the codon recognition / GTPase
activation state complex. The decoding center is
shown from the 60S side, with h44 in light blue,
h18 containing the 626 (530, E. coli numbering)
loop in dark blue, A/T-tRNA in pink,mRNA in green
and uS12 in gold. Bases 1824 and 1825 (1492 and
1493, E. coli numbering) of h44 are highlighted in
red and are shown in the flipped out positions
modeled to the codon recognition state.
(B–F) Density maps for the (B) POST state, (C) the
codon sampling state, (D) the codon recognition /
GTPase activation state, (E) the PRE classical-2
and (F) the PRE classical-1 state. The models are
kept fixed in the recognition state in all panels
for better comparison. Electron density maps
are aligned to h44 and 40S platform. For clarity,
the density around h44 and uS12 has been
segmented to show only the surface-most layer.
See also Movie S3.
change is not completed at the decoding step and still pro-
gresses from the two decoding subpopulations to the classical
PRE states (Figure 6, Movie S3).
Differences in Conformation and Interaction Pattern ofthe Ternary Complexes with the RibosomeWhen the X-ray structures of the 70S,tRNA,EF-Tu complexes
stalled by kirromycin (Schmeing et al., 2009) or GDPCP
(Voorhees et al., 2010) or the cryo-EM map of the
70S,tRNA,EF-Tu,GDP,kirromycin complex (Schuette et al.,
2009) are compared to the present 80S,Val-tRNA,eEF1A,GMPPNP cryo-EM maps by aligning the large ribosomal sub-
units, the bacterial 30S subunit is found to be rotated by �2�
and in-between the positions of the mammalian 40S subunit
of the 80S,Val-tRNA,eEF1A,GMPPNP and PRE complexes,
respectively. This is smaller than the full subunit rolling between
the mammalian POST and PRE complexes but leads to a more
open-factor-binding site of the mammalian ribosome and signif-
icant differences in the relative positions of ribosomal elements
that interact with the ternary complex. The difference in position
of the tip of the functionally important SRL, for example, is in the
range of �5 A when the 80S and 70S complexes are compared
in a common alignment of the small ribosomal subunits. From
geometric consideration some corresponding adjustment in
the ternary complex can be expected.
Indeed, when the ternary complexes are compared from the
perspective of the 40S subunit, differences in the positioning
can be detected between the bacterial codon-recognition com-
plex and the mammalian codon sampling state (Figure 4D), or
codon recognition states (Figure 4E), respectively. Positional
differences between bacterial and eukaryotic complexes mani-
fest mainly at the tRNA elbow, whereas the positions and inter-
actions of the two ends of the A/T-tRNA, i.e. the ASL and the
30-CCA-end are similar (Figures 4D and 4E). The interaction of
the 30-CCA-end with the 40S subunit occurs in the context of
an interaction with domain II of eEF1A with the shoulder region
of the 40S subunit. Consistent with their functional importance
for decoding (Voorhees and Ramakrishnan, 2013), these interac-
tions appear highly conserved from bacteria to mammals. Thus,
similar to the changes between the codon sampling and codon
recognition states of the 80S,Val-tRNA,eEF1A,GMPPNP com-
plex, the interaction of domain II with the shoulder of the 40S
subunit serves as anchor point (Figure 5A).
However, the presented mammalian model of the ternary
complex shows a subtle change in the relative orientations of
eEF1A and tRNA. Compared to the ribosome-bound Thermus
thermophilus ternary complex (Voorhees et al., 2010) the T-arm
ismoved away from the core of the protein by 4–5 A (Figure S5A).
In light of the limited resolution of our study this could be re-
garded as not meaningful. However, a movement in this range
can be predicted from a structural comparison of EF-Tu (Voo-
rhees et al., 2010) and aEF1A (Kobayashi et al., 2012). Due
to sequence variation the interaction surface of domain III of
aEF1A is grown out andwould clash with the bacterial tRNA (Fig-
ure S5A). Tentative interactions of the mammalian ternary com-
plex are formed by the loops around His349/Pro350 (human
numbering) and Asp428/Met429 (human numbering) with the
tRNA around positions 52 and 63/64, respectively (Figure S5A).
Thus, our structural findings are corroborated by evolutionary
changes of EF1A, which apparently adapt the ternary complex
to a changed ribosomal environment.
Moreover, the different position of the ternary complex in the
two states of the mammalian decoding complex and in the bac-
terial complex leads to a differential interaction pattern with the
ribosome. For the mammalian codon sampling state there is a
contact between between uS12 of the 40S shoulder and domain
III of eEF1A nearby Pro409 (human numbering) (Figure 5D) facil-
itated by a eukaryotic-specific rearrangement of the C-terminal
tail of uS12 (rpS23). In the mammalian codon recognition state,
this specific contact and also the nearby evolutionary conserved
contact between nucleotide 68 of the A/T-tRNA and protein
uS12 is less obvious than for the mammalian codon sampling
Cell 158, 121–131, July 3, 2014 ª2014 Elsevier Inc. 127
state (Figure 5D, right) and can be observed only at lower con-
tour level.
Most interestingly, the changed position of the elbow of the
eukaryotic A/T-tRNA is stabilized by specific interactions with
the 60S subunit (Figure 5C). In the mammalian codon sampling
state the T-loop (near position 62/63) and D-loop (near position
17) appear to interact with the prominent SRL and helix H89,
respectively (Figure 5C, left). Direct interaction of the A/T-tRNA
with 28S rRNA (outside of the stalk base) constitutes a surprising
difference between bacterial and eukaryotic systems. These
unique eukaryotic-specific contacts could stabilize binding of
the A/T-tRNA and make the reconstruction of the initial codon
sampling state possible. For the mammalian codon recogni-
tion/GTPase activation state there is a complex interaction
pattern between the SRL and eEF1A (Figure 5B, right). It is not
entirely clear whether the contact between the apical loop of
the SRL and the ternary complex directly involves the T-loop of
the A/T-tRNA as well or occurs indirectly via the eEF1A region
around Gln431, which is adjacent to the A/T-tRNA. In addition,
the cryo-EM density shows several contact regions between
the SRL and eEF1A. Onemay involve the P loop of eEF1A around
Asp17 and/or His95 from the switch II region (Figure 5B, right, d)
and a second one Arg69 of the switch I region (Figure 5B, right,
c). Interestingly, also eukaryotic-specific elements participate in
the interaction network, i.e. the helical insertion of eEF1A near
Gly121 and Gly131, respectively (Figure 5B, right, e and f).
The conformation of the switch I region differs between the
bacterial and the eukaryotic/archaeal systems. We note a helical
insertion next to the switch I region that is specific for eukaryotes
and archae. When the G domains of yeast eEF1A in complex
with the catalytic C terminus of its nucleotide exchange factor
eEF1Ba (Andersen et al., 2000) and aEF1A,GTP (Kobayashi
et al., 2012) are compared, this helical insertion is found to
be rotated by �90�, suggesting that it is part of an extended
switch I subdomain (Figure S5B). Interestingly, density for this
expanded sequence suggests an interaction with h14 of 18S
rRNA in the mammalian codon sampling state (Figure 5A,
left, d). In bacteria, h14 has been proposed as a contact site
for the switch I region of the translational GTPases EF-G (Connell
et al., 2007), EF-Tu (Schuette et al., 2009; Villa et al., 2009), and
EF4 (Connell et al., 2008). In X-ray maps of ribosome-bound
EF-Tu, this interaction has not been observed directly, but the
importance of the h8/h14 region for decoding has been demon-
strated (Fagan et al., 2013).
DISCUSSION
Conformational Modes of theMammalian 80SRibosomeThe ribosome is a dynamic macromolecular machine, in which
the elongation steps of translation are facilitated by large-scale
conformational changes of the ribosome and its ligands. As out-
lined by the metastable energy landscape view (Munro et al.,
2009), the ribosome is rather a stochastic Brownian machine
than a mechanical one. The spontaneous nature of confor-
mational modes causes intrinsic conformational heterogeneity,
even for compositionally and functionally defined ribosomal
complexes. Thus, rather than displaying a single, unique struc-
ture, dynamic heterogeneous ensembles are exhibited that
128 Cell 158, 121–131, July 3, 2014 ª2014 Elsevier Inc.
reflect an exchange between distinct, functionally relevant struc-
tural configurations. A prominent example is the PRE state of the
bacterial 70S ribosome (for review seeMunro et al., 2009), where
spontaneous intersubunit rotation and the fluctuation between
classical and hybrid tRNA configurations have been well
documented. Bacterial and eukaryotic ribosomes differ in their
preferential PRE state: the bacterial 70S ribosome is found pre-
dominantly in the classical nonrotated conformation. In contrast,
vacant yeast 80S ribosomes (Ben-Shem et al., 2011; Spahn
et al., 2004a) or the mammalian 80S PRE complex have been
found to prefer the rotated intersubunit state, unless an excess
of deacylated tRNA or the antibiotic cycloheximide shifts the
landscape toward the classical, nonrotated state (Budkevich
et al., 2011). It follows that functional differences between the
translational apparatus from different kingdoms or species are
not necessarily caused by gross differences in binding sites or
conformation. More subtle changes to the energy landscape
may lead to differences in the distributions of states.
Here, we have determined the cryo-EMmap of themammalian
POST state (Figure 1A). Whereas the mammalian PRE complex
coexists in at least four structurally distinct and interchangeable
substates (Budkevich et al., 2011), intrinsic large-scale con-
formational changes such as the intersubunit rotation within
the POST complex have not been found. Thus, the POST state
may be distinguished by a deep minimum of energy/enthalpy
in the energy landscape of the elongating ribosome to compen-
sate for the relatively low entropy.
Surprisingly, the POST state deviates with respect to overall
conformation not only from the rotated PRE states but also
from the classical PRE states because a novel conformational
mode, termed subunit rolling, exists for mammalian 80S ribo-
somes (Figures 3 and S3 and Movie S1). Rolling converts the
POST to the classical PRE state subunit configurations and oc-
curs largely during the accommodation step of A-site occupation
(Figure 7). Back-rolling occurs in the opposite direction during
translocation from the PRE to the POST state; however, as trans-
location involves large ratchet-like intersubunit rearrangements
including the rotation of 40S subunit relative to 60S subunit
and swivel-like rotation of the small subunit head, back-rolling
may happen not as a stand-alone step, but in combination
with intersubunit rotation/back-rotation movements (Figure 7).
It follows that the conformational landscape of the mammalian
80S ribosome is more complex than the prokaryotic one with
three overall types of subunit configuration: the classical PRE,
the rotated PRE and the POST conformations. The underlying
basis for this new conformational degree of freedom may relate
to the presence of additional, dynamic intersubunit bridges in the
eukaryotic system (Figure S3D). Moreover, bound ligands can
influence the preferred conformation and can lead to additional
intermediate conformational states. As a consequence of sub-
unit rolling, the distinction between PRE and POST states is
much more pronounced in the mammalian system, which also
has implications for the exact nature of classical tRNA-binding
sites (Figure S2).
The Mammalian Decoding ComplexWe have shown here that the mammalian decoding complex
with the ternary complex being trapped in the A/T-state by
Figure 7. The Mammalian Elongation Circle Features Unique
Motions of the 40S Subunit
Cartoon representation of the eukaryotic elongation circle highlighting the
individual subunit motions necessary to convert one functional intermediate
state to the next. Movements of the small ribosomal subunit shared with the
prokaryotic system are noted in black, while eukaryotic-specific intersubunit
movements, i.e., rolling and back-rolling, are noted in red.
GMPPNP also exhibits subtle intrinsic structural dynamics as it
coexists in (at least) two substates. Because of the local config-
uration of the ribosomal decoding center (Figure 6) and the differ-
ential interaction pattern of eEF1A with the SRL (Figures 5B and
5C), we interpret them as states representing codon sampling
and codon recognition /GTPase activation, respectively. Both
substates are present essentially in a nonrotated, nonrolled
POST-like configuration. As the A-site is more open than in the
bacterial 70S complex, the binding site for the ternary complex
is altered between the bacterial and eukaryotic domains of life.
Indeed, we can observe conserved as well as divergent features
in the conformation/structure of the ternary complex itself and in
the interaction patterns of ribosomal ligandswith the ribosome to
account for this difference.
The interactions of the shoulder region of the 40S subunit with
two evolutionary conserved loops of domain II of eEF1A and
the 30-CCA-end of the aa-tRNA acts as an evolutionarily pre-
served anchor (Figures 4D, 4E, and 5A). On the other hand, local
changes in the ribosome exist in the elbow position of tRNA and
the exact location of eEF1A (Figure S5A). For example, the di-
vergent structure of the C-terminal end of uS12 (rpS23) is
seen to interact with eEF1A predominantly in the mammalian
codon sampling state (Figure 5D). Also, an insertion sequence
in eEF1A domain III dictates a change in relative orientation to
the T-loop/stem of the aa-tRNA (Figure S5A). Apparently, the
ternary complex evolved to interact with the altered configura-
tion of factor-binding sites between bacteria and mammals.
Most unexpected is the observed interaction of the A/T-tRNA
elbow with the apical loop of the highly conserved SRL and helix
H89 in the mammalian complex (Figure 5C). In the bacterial
system the tRNA elbow interacts only with the mobile stalk
base region of 23S rRNA. This indicates a more rigid and tighter
binding state for the tRNA elbow in the mammalian system.
The Mammalian Decoding PathwaySelection of the cognate tRNA has to proceed with optimal
speed and accuracy. This is achieved by a complex, multistep
pathway involving an initial selection step and a kinetic proof-
reading step (Rodnina and Wintermeyer, 2001; Geggier et al.,
2010). Crucial for tRNA selection is the codon recognition step
in the decoding center, and in particular the stabilization
of codon-anticodon interaction by A-minor interactions with
A1492/A1493 of 16S rRNA (A1824/A1825 in human) in the flip-
ped-out conformation (Ogle et al., 2002). Therefore, enforcing
Watson-Crick geometry of the codon-anticodon duplex pro-
vides discrimination energy between the cognate and near-
cognate interactions. Conformational changes in the ribosomes
decoding center in turn provide the signal for activation of GTP
hydrolysis by EF-Tu/eEF1A about 80 A away. For the bacterial
system, cryo-EM (Schuette et al., 2009; Villa et al., 2009) and
finally X-ray crystallography (Schmeing et al., 2009; Voorhees
et al., 2010) have revealed a series of coupled conformational
changes that start at the decoding center and are transferred
via domain closure of the 30S subunit and a distortion of the
tRNA to EF-Tu, inducing the activated conformation. Less clear
is the preceding step, because direct structural information
about the codon-sampling state is lacking.
For the mammalian sytem, however, we have now obtained
the cryo-EMmap of a substate of the GMPPNP-stalled mamma-
lian decoding complex that shows all hallmark features of a
codon sampling state (Figures 4 and 5). The ASL of the ternary
complex is located in the decoding center allowing the formation
of the codon-anticodon duplex. However, the appearance of the
ribosomal decoding center indicates that the stabilizing A-minor
interactions with A1824/A1825 of 18S rRNA do not yet occur, in
contrast to the codon recognition state (Figures 6C and 6D).
Interestingly, the conformation of the A/T-tRNA is in a bent
conformation also for the codon sampling state, likely facilitating
efficient monitoring of the codon. In the mammalian system,
codon recognition is apparently not essential for keeping the
tRNA in the distorted conformation.
Comparison of both subpopulations of the GMPPNP-stalled
decoding complex suggests a model for the mammalian decod-
ing pathway with similarities, but also surprising distinctions to
the generally accepted model for the bacterial system (Voorhees
and Ramakrishnan, 2013). As in the bacterial system, stabiliza-
tion of the codon-anticodon duplex by A-minor interactions
with A1824/A1825 of 18S rRNA and long-range signaling of
this event to the GTPase center of eEF1A are key for decoding.
However, in eukaryotes codon recognition results in a subtle
movement of the ASL deeper into the decoding cleft (Movie
S2). With the interaction between the two evolutionary
conserved 258GIGTV and 289VKS/312VKN loops of eEF1A domain
II and the shoulder region of the 40S subunit serving as an an-
chor, pulling on the ASL leads in first approximation to a rotation
of the ternary complex as a whole. Due to a lever arm effect,
movements are largest at the elbow region of the tRNA and the
Gdomain of eEF1A.We do not exclude the presence of concom-
itant conformational changes in the ternary complex such as a
Cell 158, 121–131, July 3, 2014 ª2014 Elsevier Inc. 129
smaller rotation of the G domain of the factor relative to domains
II and III that has been observed in the bacterial system (Voo-
rhees et al., 2010). As a result of the rotational movement of
the ternary complex, the G domain of eEF1A becomes posi-
tioned at the SRL by a seesaw-like mechanism. The SRL has
been suggested to play a prominent role by facilitating a rear-
rangement of a catalytically important His84 (E. coli nomencla-
ture) within the switch II regions of translational GTPase factors
(Connell et al., 2007; Schuette et al., 2009; Voorhees et al.,
2010). Thus, the codon-recognition-dependent movement of
eEF1A may lead to further conformational changes within the
G-nucleotide-binding site resulting in the GTPase activated
state.
CONCLUSIONS
We have presented here cryo-EM maps of the mammalian 80S
ribosome POST and decoding complexes at subnanometer res-
olution. The maps reveal mechanistic differences between the
mammalian and the bacterial elongation cycle. Most surp-
risingly, a unique rolling of the ribosomal 40S subunit by �6�
around the upper region of the h44 occurs during the transition
from the POST to the PRE state. Accordingly, there must be
changes during the tRNA selection step and indeed we can
describe some prominent differences in the conformation of
the ribosome-bound aa-tRNA,eEF1A,GMPPNP ternary com-
plex as well as its interaction pattern with the ribosome. Evolu-
tionary changes in key steps of elongation demonstrate that in
structural terms the bacterial and the mammalian elongation cy-
cle are less similar than previously thought. Further studies will
be needed to understand whether and how these structural
changes translate into changes of mechanisms or functions in
order to achieve optimal translation or translational control in
the eukaryotic environment.
EXPERIMENTAL PROCEDURES
Sample Preparation
Reassociated 80S ribosomes from rabbit liver, free of endogenous tRNAs and
mRNAs, were prepared according to (Bommer et al., 1997). The POST 80S
complex bearing deacylated tRNAPhe in the E site and N-acetyl-Lys-tRNALys3
in the P-site was prepared by addition of elongation factor 2 (eEF2) in the
presence of 200 mM GTP to PRE complex (0.8 mM) programmed with MFK-
mRNA (Budkevich et al., 2008). The occupancy of N-acylated Lys-tRNALys3
was approximately 0.66 per 80S ribosome and more than 90% of the bound
tRNA was reactive with puromycin, indicating a nearly quantitative transloca-
tion and P-site location. In order to prepare the decoding complex, 80S ribo-
somes programmed with MFV-mRNA and occupied by Ac[3H]Phe-tRNAPhe
were incubated with preformed ternary complex eEF1A,GMPPNP,[14C]Val-
tRNAVal in the presence of 400 mM GMPPNP (Budkevich et al., 2008).
Electron Microscopy and Image Processing
The 80S complexes were diluted to a final concentration of 30 nM and flash-
frozen in liquid ethane and images were recorded under low-dose conditions
using an FEI Tecnai G2 Polara operating at 300 kV and a nominal magnification
of 39,000. The resulting micrographs were digitized on a drum-scanner (Hei-
delberg) with a pixel size of 1.26 A on the object scale. Multiparticle refinement
using SPIDER (Frank et al., 1996) was carried out as described previously
(Budkevich et al., 2011; Loerke et al., 2010; Penczek et al., 2006) in order to
overcome sample heterogeneity caused by substochiometric binding of the
ligands and inhomogeneity in the preparation of themammalian 80S ribosome.
130 Cell 158, 121–131, July 3, 2014 ª2014 Elsevier Inc.
Final reconstructions were calculated with SPARX (Hohn et al., 2007). The final
resolution for all maps was estimated using the 0.5 cutoff criteria from the
Fourier shell correlation curves. For further details see Extended Experimental
Procedures.
Modeling
A secondary structure map for rabbit 80S ribosomes is not available, so
the human ribosome as target for the modeling was chosen. Initial structural
models of the human ribosomal RNAswere built following a homologymodeling
approach (Tung and Sanbonmatsu, 2004). For homology modeling of the ribo-
somal proteins we used Prime (from the Schrodinger suite) (Jacobson et al.,
2002; Jacobson et al., 2004). Initial structural models of the 80S ribosome
were fit to the cryo-EM map of the 80S POST complex using MDfit (Ratje
etal., 2010)whichallowsflexibledockingwhilemaintainingstereochemistrypre-
sent in the initialmodel. Simulationswereperformedatstructure-basedpotential
temperature of 20. EMweightwas adjusted to the number of atoms in themodel.
ACCESSION NUMBERS
The European Molecular Biology Laboratory-European Bioinformatics Insti-
tute (Cambridge, UK) 3D-EM data base accession numbers for the electron
density maps of the mammalian POST, PRE (classical-1/2), and decoding
(codon sampling and codon recognition/ GTPase activation) complexes re-
ported in this paper are EMD-2620, EMD-2621, EMD-2622, EMD-2623 and
EMD-2624, respectively. The Protein Data Bank accession numbers for the
human models for 40S and 60S subunits are 4cxc (40S), 4cxd (60S ribosomal
proteins), 4cxe (60S rRNA), and 4cxb (ligands) and for the ternary complexes
are 4cxg (initial codon sampling state) and 4cxh (codon recognition state).
SUPPLEMENTAL INFORMATION
Supplemental Information includes Extended Experimental Procedures, five
figures, one table, and three movies and can be found with this article online
at http://dx.doi.org/10.1016/j.cell.2014.04.044.
AUTHOR CONTRIBUTION
T.V.B. prepared the 80S-POST and decoding complexes. T.V.B., J.G. and
T.M. collected the cryo-EM data. T.V.B., J.G., J.L., and C.M.T.S. carried out
the image processing. E.B., J.L., D.J.F.R., J.I., P.W.H., C.-S.T., K.Y.S. and
C.M.T.S. carried out the modeling of the human 80S ribosomes. T.V.B.,
E.B., J.L., K.H.N. and C.M.T.S. discussed the results and wrote the paper.
ACKNOWLEDGMENTS
We thankDr. Scott Blanchard for helpful discussion. Thepresentworkwas sup-
ported by grants from the Deutsche Forschungsgemeinschaft DFG (SFB 740
to C.M.T.S., P.W.H., and T.M.; 436 UKR 113/64/1-1 to T.B. and HI 1502 to
P.W.H.), HSFP and Senatsverwaltung fur Wissenschaft, Forschung und Kultur
Berlin (UltraStructureNetwork, Anwenderzentrum). K.Y.S. was supported by
HFSP, NIH Grant R01-GM072686 and Los Alamos Institutional Computing.
We acknowledge the use of computational resources supplied by the North-
German Supercomputing Alliance (HLRN; project beb00001 to C.M.T.S and
bec00085 to P.W.H.).
Received: October 1, 2013
Revised: February 24, 2014
Accepted: April 18, 2014
Published: July 3, 2014
REFERENCES
Andersen, G.R., Pedersen, L., Valente, L., Chatterjee, I., Kinzy, T.G., Kjeldg-
aard, M., and Nyborg, J. (2000). Structural basis for nucleotide exchange
and competition with tRNA in the yeast elongation factor complex
eEF1A:eEF1Balpha. Mol. Cell 6, 1261–1266.
Andersen, C.B., Becker, T., Blau,M., Anand,M., Halic, M., Balar, B., Mielke, T.,
Boesen, T., Pedersen, J.S., Spahn, C.M., et al. (2006). Structure of eEF3 and
the mechanism of transfer RNA release from the E-site. Nature 443, 663–668.
Anger, A.M., Armache, J.P., Berninghausen, O., Habeck, M., Subklewe, M.,
Wilson, D.N., and Beckmann, R. (2013). Structures of the human and
Drosophila 80S ribosome. Nature 497, 80–85.
Ban, N., Beckmann, R., Cate, J.H., Dinman, J.D., Dragon, F., Ellis, S.R., Lafon-
taine, D.L., Lindahl, L., Liljas, A., Lipton, J.M., et al. (2014). A new system for
naming ribosomal proteins. Curr. Opin. Struct. Biol. 24, 165–169.
Ben-Shem, A., Garreau de Loubresse, N., Melnikov, S., Jenner, L., Yusupova,
G., and Yusupov, M. (2011). The structure of the eukaryotic ribosome at 3.0 A
resolution. Science 334, 1524–1529.
Bommer, U., Burkhardt, N., Junemann, R., Spahn, C.M.T., Triana-Alonso, F.,
and Nierhaus, K.H. (1997). Ribosomes and polysomes. In Subcellular Fraction-
ation: A practical Approach, J. Graham and D. Rickwood, eds. (Washington,
DC: IRL Press), pp. 271–301.
Budkevich, T.V., El’skaya, A.V., and Nierhaus, K.H. (2008). Features of 80S
mammalian ribosome and its subunits. Nucleic Acids Res. 36, 4736–4744.
Budkevich, T., Giesebrecht, J., Altman, R.B., Munro, J.B.,Mielke, T., Nierhaus,
K.H., Blanchard, S.C., and Spahn, C.M. (2011). Structure and dynamics of the
mammalian ribosomal pretranslocation complex. Mol. Cell 44, 214–224.
Connell, S.R., Takemoto, C., Wilson, D.N., Wang, H., Murayama, K., Terada,
T., Shirouzu, M., Rost, M., Schuler, M., Giesebrecht, J., et al. (2007). Structural
basis for interaction of the ribosome with the switch regions of GTP-bound
elongation factors. Mol. Cell 25, 751–764.
Connell, S.R., Topf, M., Qin, Y., Wilson, D.N., Mielke, T., Fucini, P., Nierhaus,
K.H., and Spahn, C.M. (2008). A new tRNA intermediate revealed on the
ribosome during EF4-mediated back-translocation. Nat. Struct. Mol. Biol.
15, 910–915.
Fagan, C.E., Dunkle, J.A., Maehigashi, T., Dang, M.N., Devaraj, A., Miles, S.J.,
Qin, D., Fredrick, K., and Dunham, C.M. (2013). Reorganization of an intersu-
bunit bridge induced by disparate 16S ribosomal ambiguity mutations mimics
an EF-Tu-bound state. Proc. Natl. Acad. Sci. USA 110, 9716–9721.
Frank, J., and Spahn, C.M. (2006). The ribosome and themechanism of protein
synthesis. Rep. Prog. Phys. 69, 1383–1417.
Frank, J., Radermacher, M., Penczek, P., Zhu, J., Li, Y., Ladjadj, M., and Leith,
A. (1996). SPIDER and WEB: processing and visualization of images in 3D
electron microscopy and related fields. J. Struct. Biol. 116, 190–199.
Geggier, P., Dave, R., Feldman, M.B., Terry, D.S., Altman, R.B., Munro, J.B.,
and Blanchard, S.C. (2010). Conformational sampling of aminoacyl-tRNA dur-
ing selection on the bacterial ribosome. J. Mol. Biol. 399, 576–595.
Hohn, M., Tang, G., Goodyear, G., Baldwin, P.R., Huang, Z., Penczek, P.A.,
Yang, C., Glaeser, R.M., Adams, P.D., and Ludtke, S.J. (2007). SPARX, a
new environment for Cryo-EM image processing. J. Struct. Biol. 157, 47–55.
Ingolia, N.T., Lareau, L.F., and Weissman, J.S. (2011). Ribosome profiling of
mouse embryonic stem cells reveals the complexity and dynamics of mamma-
lian proteomes. Cell 147, 789–802.
Jacobson, M.P., Friesner, R.A., Xiang, Z., and Honig, B. (2002). On the role
of the crystal environment in determining protein side-chain conformations.
J. Mol. Biol. 320, 597–608.
Jacobson, M.P., Pincus, D.L., Rapp, C.S., Day, T.J., Honig, B., Shaw, D.E.,
and Friesner, R.A. (2004). A hierarchical approach to all-atom protein loop
prediction. Proteins 55, 351–367.
Jenner, L.B., Demeshkina, N., Yusupova, G., and Yusupov, M. (2010). Struc-
tural aspects of messenger RNA reading frame maintenance by the ribosome.
Nat. Struct. Mol. Biol. 17, 555–560.
Klinge, S., Voigts-Hoffmann, F., Leibundgut, M., Arpagaus, S., and Ban, N.
(2011). Crystal structure of the eukaryotic 60S ribosomal subunit in complex
with initiation factor 6. Science 334, 941–948.
Kobayashi, K., Saito, K., Ishitani, R., Ito, K., and Nureki, O. (2012). Structural
basis for translation termination by archaeal RF1 and GTP-bound EF1a
complex. Nucleic Acids Res. 40, 9319–9328.
Loerke, J., Giesebrecht, J., and Spahn, C.M. (2010). Multiparticle cryo-EM of
ribosomes. Methods Enzymol. 483, 161–177.
Melnikov, S., Ben-Shem, A., Garreau de Loubresse, N., Jenner, L., Yusupova,
G., and Yusupov, M. (2012). One core, two shells: bacterial and eukaryotic
ribosomes. Nat. Struct. Mol. Biol. 19, 560–567.
Munro, J.B., Sanbonmatsu, K.Y., Spahn, C.M., and Blanchard, S.C. (2009).
Navigating the ribosome’s metastable energy landscape. Trends Biochem.
Sci. 34, 390–400.
Ogle, J.M., Murphy, F.V., Tarry, M.J., and Ramakrishnan, V. (2002). Selection
of tRNA by the ribosome requires a transition from an open to a closed form.
Cell 111, 721–732.
Penczek, P.A., Yang, C., Frank, J., and Spahn, C.M. (2006). Estimation of vari-
ance in single-particle reconstruction using the bootstrap technique. J. Struct.
Biol. 154, 168–183.
Rabl, J., Leibundgut, M., Ataide, S.F., Haag, A., and Ban, N. (2011). Crystal
structure of the eukaryotic 40S ribosomal subunit in complex with initiation
factor 1. Science 331, 730–736.
Ratje, A.H., Loerke, J., Mikolajka, A., Brunner, M., Hildebrand, P.W., Starosta,
A.L., Donhofer, A., Connell, S.R., Fucini, P., Mielke, T., et al. (2010). Head
swivel on the ribosome facilitates translocation by means of intra-subunit
tRNA hybrid sites. Nature 468, 713–716.
Rodnina, M.V., and Wintermeyer, W. (2001). Fidelity of aminoacyl-tRNA selec-
tion on the ribosome: kinetic and structural mechanisms. Annu. Rev. Biochem.
70, 415–435.
Sanbonmatsu, K.Y., Joseph, S., and Tung, C.S. (2005). Simulating movement
of tRNA into the ribosome during decoding. Proc. Natl. Acad. Sci. USA 102,
15854–15859.
Schilling-Bartetzko, S., Bartetzko, A., and Nierhaus, K.H. (1992). Kinetic and
thermodynamic parameters for tRNA binding to the ribosome and for the
translocation reaction. J. Biol. Chem. 267, 4703–4712.
Schmeing, T.M., Voorhees, R.M., Kelley, A.C., Gao, Y.G., Murphy, F.V., 4th,
Weir, J.R., and Ramakrishnan, V. (2009). The crystal structure of the ribosome
bound to EF-Tu and aminoacyl-tRNA. Science 326, 688–694.
Schuette, J.C., Murphy, F.V., 4th, Kelley, A.C., Weir, J.R., Giesebrecht, J.,
Connell, S.R., Loerke, J., Mielke, T., Zhang, W., Penczek, P.A., et al. (2009).
GTPase activation of elongation factor EF-Tu by the ribosome during decod-
ing. EMBO J. 28, 755–765.
Spahn, C.M., Jan, E., Mulder, A., Grassucci, R.A., Sarnow, P., and Frank, J.
(2004a). Cryo-EM visualization of a viral internal ribosome entry site bound
to human ribosomes: the IRES functions as an RNA-based translation factor.
Cell 118, 465–475.
Spahn, C.M., Gomez-Lorenzo, M.G., Grassucci, R.A., Jørgensen, R., Ander-
sen, G.R., Beckmann, R., Penczek, P.A., Ballesta, J.P., and Frank, J.
(2004b). Domain movements of elongation factor eEF2 and the eukaryotic
80S ribosome facilitate tRNA translocation. EMBO J. 23, 1008–1019.
Steitz, J.A. (1969). Nucleotide sequences of the ribosomal binding sites of
bacteriophage R17 RNA. Cold Spring Harb. Symp. Quant. Biol. 34, 621–630.
Tung, C.S., and Sanbonmatsu, K.Y. (2004). Atomic model of the Thermus ther-
mophilus 70S ribosome developed in silico. Biophys. J. 87, 2714–2722.
Villa, E., Sengupta, J., Trabuco, L.G., LeBarron, J., Baxter, W.T., Shaikh, T.R.,
Grassucci, R.A., Nissen, P., Ehrenberg, M., Schulten, K., and Frank, J. (2009).
Ribosome-induced changes in elongation factor Tu conformation control GTP
hydrolysis. Proc. Natl. Acad. Sci. USA 106, 1063–1068.
Voorhees, R.M., and Ramakrishnan, V. (2013). Structural basis of the transla-
tional elongation cycle. Annu. Rev. Biochem. 82, 203–236.
Voorhees, R.M., Schmeing, T.M., Kelley, A.C., and Ramakrishnan, V. (2010).
The mechanism for activation of GTP hydrolysis on the ribosome. Science
330, 835–838.
Wilson, D.N. (2009). The A-Z of bacterial translation inhibitors. Crit. Rev.
Biochem. Mol. Biol. 44, 393–433.
Cell 158, 121–131, July 3, 2014 ª2014 Elsevier Inc. 131
Supplemental Information
EXTENDED EXPERIMENTAL PROCEDURES
Sample PreparationReassociated 80S ribosomes from rabbit liver, free of endogenous tRNAs and mRNAs, were prepared according to (Bommer et al.,
1997) with slight modifications. In order to wash endogenous RNases from polysomes, the postmitochondrial supernatant was
centrifuged through a 1M sucrose cushion, which contained 0.5 M KCl. Reassociation of 40S and 60S subunits was confirmed by
analytical centrifugation in SW40 rotor (18,000 rpm, 17 hr). The ribosome concentrations were calculated assuming the following
ratios: 60 pmol/A260 for 40S subunits, 30 pmol/A260 for 60S subunits and 20 pmol/A260 for 80S ribosomes. 80S particles were pro-
grammedwith the heteropolymeric MFK-mRNA for POST, andMFV-mRNA for decoding complex preparedwith run-off transcription
according to (Triana-Alonso et al., 1995). Bovine liver tRNAPhe, tRNALys3 and tRNAVal were purified from tRNAbulk preparations by us-
ing ion-exchange DEAE and reverse phase Hypersil 5 C4 HPLC chromatography. N-acetyl-[14C]Lys-tRNALys3 , N-acetyl-[3H]Phe-
tRNAPhe, and [14C]Val-tRNAVal were purified by reverse-phase HPLC on a Nucleosil 300-5 C4 column using a methanol gradient
as described (Cayama et al., 2000).
Elongation factors eEF1A and eEF2 from rabbit liver were isolated using a combination of gel-filtration and ion-exchange chroma-
tography as described (Kemper and Merrick, 1979; Shalak et al., 1997).
The extent of tRNA bindingwas determined by nitrocellulose filtration and the specificity of tRNA binding by the puromycin reaction
(0.8 mM). All incubations were performed in polyamine buffer containing 20 mM HEPES–KOH (pH 7.5 at 4�C), 5 mMMgCl2, 100 mM
KCl, 0.6 mM spermine, 0.8 mM spermidine and 6 mM 2-mercaptoethanol (El’skaya et al., 1997).
Electron Microscopy and Image ProcessingReassociated 80S complexes were diluted to a final concentration of 30 nM and flash-frozen in liquid ethane on carbon-coated
Quantifoil grids (Quantifoil, Germany) using a Vitrobot (FEI) device. Images were recorded under low-dose conditions on film (Kodak
SO-163) using an FEI Tecnai G2 Polara operating at 300 kV and a nominal magnification of 39,000. The resulting micrographs were
digitized on a drum-scanner (Heidelberg) with a pixel size of 1.26 A on the object scale. After visual inspection and evaluation of
micrographs according to their power spectra, particles were preselected using Signature (Chen and Grigorieff, 2007), boxed out
and decimated. This and all further image processing steps were carried out using SPIDER (Frank et al., 1996), final reconstructions
were calculated with SPARX (Hohn et al., 2007). In order to overcome sample heterogeneity caused by substochiometric binding of
the ligands and inhomogeneity in the preparation of the mammalian 80S ribosome, multiparticle refinement was carried out as
described previously (Budkevich et al., 2011; Loerke et al., 2010; Penczek et al., 2006). A reconstruction of the vacant 80S ribosome
from rabbit was introduced as a seed structure to increase the number of references until stable subpopulations were obtained.
The 80S POST complex (initial data set is 826 micrographs with 1,072,632 particle images) was analyzed and sorted according to
multi-particle alignment procedures (Figure S1). During this procedure about 400,000 particle images were discarded. Only about
35% (236,113) of the remaining ribosomal complexes constituted the subpopulation corresponding to the desired 80S POST com-
plex and were used for final reconstruction (Figure S1). In addition, our unsupervised classification procedure revealed the presence
of two subpopulations representing PRE complexes. Themap of the larger PRE subpopulation (29%of the particle images) exhibited
density for tRNAs in classical A-, P- and E-sites, whereas the second PRE complex (15% of the particle images) had tRNAs in hybrid
A/P- and P/E-states. According to our previous analysis of the 80S PRE complex (Budkevich et al., 2011) these two PRE subpop-
ulations correspond to the classical-1 and rotated-2 PRE states, respectively. Further refinement of the PRE classical-1 state yielded
three distinct structures: PRE (classical-1: 66,618 images), PRE (classical-2: 73,951 images) and an unassigned structure with frag-
mented A-site tRNA (51,149 images).
In addition to the POST and PRE-like structures, the initial data set contained a fourth subpopulation of particle images (143,150
images) that exhibited ligand density in the factor-binding site together with density for P/P- and E/E-tRNAs. By its appearance this
density can be identified to correspond to elongation factor eEF2 (Figure S1). It is remarkable that eEF2 is found bound to a subset of
ribosomes without being stalled by an antibiotic or a nonhydrolyzable GTP analog. The significant presence of PRE complexes and
translocation intermediates within the posttranslocational sample cannot be explained by inefficient translocation, because the pu-
romycin reaction indicates nearly quantitative translocation. However, the biochemical analysis and the cryo-EM results can be
reconciled by taking into account the possibility of a reverse translocation reaction. Spontaneous, reverse translocation has been
found in bacterial system and can be driven by an excess of deacylated tRNA in in vitro assays (Shoji et al., 2006). The possibility
of a reverse translocation was also suggested for the eukaryotic system (Nilsson and Nygard, 1992). The presence of eEF2 can
lead to the recurring translocation. It seems that translocation and reverse translocation result in equilibrium of PRE and POST com-
plexes as visualized by cryo-EM. eEF2-dependent translocation can occur also during the incubation with puromycin leading to a
nearly quantitative puromycin reaction.
For the 80S decoding complex, the initial data set (230 micrographs, 626,562 particle images) separated into two major
populations during several rounds of the multi-particle refinement. The first subpopulation, reconstructed from 247,633 particle
images, represents the decoding complex with two classically oriented tRNAs in P- and E-sites and the ternary complex aa-
tRNA,eEF1A,GMPPNP stalled in the A/T-position. The second subpopulation, reconstructed from 210,574 particle images, consists
of PRE 80S ribosomes bearing three tRNA molecules and was described in (Budkevich et al., 2011). For further processing, only the
Cell 158, 121–131, July 3, 2014 ª2014 Elsevier Inc. S1
first of these two subpopulations was used. After further multi-particle classification and removal of particles images not assigned to
the decoding complex, the remaining 176,675 particles split into three final subpopulations: the codon sampling complex (79,705
particle images), the codon recognition / GTPase activation complex (52,686 particle images), and a third, unassigned population
with 44,284 particle images.
The final resolution for all (POST, classical-1, classical-2, codon sampling and codon recognition / GTPase activation) maps was
estimated using the 0.5 cutoff criteria from the Fourier shell correlation curves (Figure S1B) and the maps were filtered accordingly.
Interpretation and ModelingAs a starting point, models derived from the X-ray structures of S. cerevisiae 80S (Ben-Shem et al., 2010; PDB IDs 3O2Z, 3O58; Ben-
Shem et al., 2011; PDB IDs 3U5B-I), T. thermophila 40S-eIF1 complex (Rabl et al., 2011; PDB ID 2XZN) and T. thermophila 60S-eIF6
(Klinge et al., 2011; PDB IDs 4A17, 4A19) were fit as rigid-bodies into the cryo-EMmaps usingChimera (Pettersen et al., 2004). Crystal
structures of prokaryotic tRNAs (Voorhees et al., 2010; PDB ID 2WDG) were docked into segmented densities corresponding to the
tRNAs. For A/T-tRNA, we created hybrid structures from PDB IDs 2XQD (27-43) and 1TTT (1-26; 44-76) by fitting each fragment inde-
pendently as a rigid-body into the decoding state maps. RNA geometry at the junction sites was then restored using RCrane (Keating
and Pyle, 2012) and CNS 1.3 (Brunger, 2007).
As a secondary structure map for rabbit 80S ribosomes is not available, we chose the human ribosome as target for our modeling.
Importantly, on an overall structural level rabbit and human ribosomes are very similar (Budkevich et al., 2011; Spahn et al., 2004b).
Furthermore, sequence alignment between ribosomal proteins from human and rabbit shows that the sequence identity is generally
high with the majority of the aligned proteins being identical in length and sequence (for example eS6, uS9 (rpS16), uS12 (rpS23),
eS24, eS27, uL15 (L27A), eL19, eL27). Others are identical in length but show minor differences in sequence identity (e.g., eS8
99%, eS26 87%, eL32 94%, eL39 78%). Only few are slightly different in length and sequence identity (e.g., eL6: human 288 residues
versus rabbit 291, 84%).
It is remarkable that mammalian 80S ribosomes contain only one additional protein, eL28 (L28e), compared to ribosomes from
yeast. However, protein eL28 is present in T. thermophila 60S subunits (Klinge et al., 2011). To obtain an all-atommodel highly consis-
tent with the cryo-EM reconstructions, we also employed MDfit (Whitford et al., 2011), which allows for flexible docking while main-
taining stereochemistry present in the initial model. As we did not attempt to model those parts of rRNA expansion elements and
proteins that are apparently too flexible to be observed in the map, our final model of the human 80S ribosome encompasses 33 pro-
teins from the small and 43 proteins from the large subunit and 5,634 out of 7,173 nucleotides for the rRNAs (Figure 1A, Table S1).
The initial structural models of the human ribosomal RNAs were calculated following a protocol developed for E. coli 30S (Tung
et al., 2002) and T. thermophilus 70S (Tung and Sanbonmatsu, 2004) ribosomal RNA modeling. The 4 A resolution crystal structures
(Ben-Shem et al., 2010; PDB IDs 3O2Z, 3O58) of the yeast small and large ribosomal subunits were used as templates for RNA ho-
mologymodeling. The resultingmodels were further improved using the 3 A resolution crystal structures (Ben-Shem et al., 2011; PDB
IDs 3U5B, 3U5D) of the yeast small and large ribosomal subunits. Ourmotif-based approach preserves RNA stereochemistry present
in crystallographic motifs and maintains structural integrity (e.g., bond lengths, van der Waals contacts).
For homology modeling of the ribosomal proteins we used Prime (from the Schrodinger suite) (Jacobson et al., 2002, Jacobson
et al., 2004). The crystal structures from yeast (PDB ID 3U5E) or Tetrahymena thermophila (PDB ID 2XZM) were used as templates.
Appropriate template structures were selected from these two entries based on completeness of the template structure, similarity
between target and template and fitting of the modeled structure to the electron density maps. Loops and insertions were either
modeled with Prime (Schrodinger suite) or with Superlooper (Hildebrand et al., 2009). Superlooper allows visualization and selection
of a loop conformation in the context of the whole molecule. 87% of all residues of these 76 proteins, 4,745 of 5,476 residues of
the small subunit and 6,507 of 7,429 residues of the large subunit, were modeled. The average sequence identity/similarity between
template and target protein structures is 50%/67% for the small subunit and 55%/70% for the large subunit. The sequence identity
between target and template protein structures in the small and the large subunit ranges from 22% (eL28) to 88% (eL41).
Initial structural models of the 80S ribosome were flexibly fit to the cryo-EM reconstruction of the 80S POST configuration using
MDfit (Whitford et al., 2011). Simulations used protocols similar to those in the online tutorial and to our previous studies (Ratje et al.,
2010). Forces associated with EM density were evaluated every 200 time steps. Simulations were performed at structure-based
potential temperature of 20. EMweight was adjusted to the number of atoms in the model. As in previous studies, three successive
simulations were performed: (1) MDfit using the 80S cryo-EM reconstruction with an additional Gaussian filter with a half-width of 4 A,
(2) MDfit using the 80S cryo-EM reconstruction Gaussian with an additional Gaussian filter with a half-width of 2 A, and (3) MDfit using
the 80S cryo-EM reconstruction. Each simulation used between 100,000-300,000 time steps and required �24 hr on a MAC OSX
desktop computer with eight processor cores.
Cryo-EMdensities and the atomicmodels were visualized using the programChimera (Pettersen et al., 2004), which was also used
to create all images and movies.
SUPPLEMENTAL REFERENCES
Apweiler, R., Bairoch, A., Wu, C.H., Barker, W.C., Boeckmann, B., Ferro, S., Gasteiger, E., Huang, H., Lopez, R., Magrane, M., et al. (2004). UniProt: the Universal
Protein knowledgebase. Nucleic Acids Res. 32 (Database issue), D115–D119.
S2 Cell 158, 121–131, July 3, 2014 ª2014 Elsevier Inc.
Ben-Shem, A., Jenner, L., Yusupova, G., and Yusupov, M. (2010). Crystal structure of the eukaryotic ribosome. Science 330, 1203–1209.
Brunger, A.T. (2007). Version 1.2 of the Crystallography and NMR system. Nat. Protoc. 2, 2728–2733.
Cayama, E., Yepez, A., Rotondo, F., Bandeira, E., Ferreras, A.C., and Triana-Alonso, F.J. (2000). New chromatographic and biochemical strategies for quick
preparative isolation of tRNA. Nucleic Acids Res. 28, E64.
Chen, J.Z., and Grigorieff, N. (2007). SIGNATURE: a single-particle selection system for molecular electron microscopy. J. Struct. Biol. 157, 168–173.
El’skaya, A.V., Ovcharenko, G.V., Palchevskii, S.S., Petrushenko, Z.M., Triana-Alonso, F.J., and Nierhaus, K.H. (1997). Three tRNA binding sites in rabbit liver
ribosomes and role of the intrinsic ATPase in 80S ribosomes from higher eukaryotes. Biochemistry 36, 10492–10497.
Hildebrand, P.W., Goede, A., Bauer, R.A., Gruening, B., Ismer, J., Michalsky, E., and Preissner, R. (2009). SuperLooper—a prediction server for the modeling of
loops in globular and membrane proteins. Nucleic Acids Res. 37 (Web Server issue), W571–W574.
Keating, K.S., and Pyle, A.M. (2012). RCrane: semi-automated RNA model building. Acta Crystallogr. D Biol. Crystallogr. 68, 985–995.
Kemper, W.M., and Merrick, W.C. (1979). Preparation of protein synthesis elongation factors from rabbit reticulocytes. Methods Enzymol. 60, 638–648.
Kjeldgaard, M., Nyborg, J., and Clark, B.F. (1996). The GTP binding motif: variations on a theme. FASEB J. 10, 1347–1368.
Nilsson, L., and Nygard, O. (1992). Reduced puromycin sensitivity of translocated polysomes after the addition of elongation factor 2 and non-hydrolysable GTP
analogues. FEBS Lett. 309, 89–91.
Pettersen, E.F., Goddard, T.D., Huang, C.C., Couch, G.S., Greenblatt, D.M., Meng, E.C., and Ferrin, T.E. (2004). UCSF Chimera—a visualization system for
exploratory research and analysis. J. Comput. Chem. 25, 1605–1612.
Rhodin, M.H., and Dinman, J.D. (2010). A flexible loop in yeast ribosomal protein L11 coordinates P-site tRNA binding. Nucleic Acids Res. 38, 8377–8389.
Shalak, V.F., Budkevich, T.V., Negrutski1, B.S., and El’skaia, A.V. (1997). A fast and effective method for purification of elongation factor 1 alpha from rabbit liver.
Ukr. Biokhim. Zh. 69, 104–109.
Shoji, S., Walker, S.E., and Fredrick, K. (2006). Reverse translocation of tRNA in the ribosome. Mol. Cell 24, 931–942.
Sievers, F., Wilm, A., Dineen, D., Gibson, T.J., Karplus, K., Li, W., Lopez, R., McWilliam, H., Remmert, M., Soding, J., et al. (2011). Fast, scalable generation of
high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 7, 539.
Spahn, C.M., Beckmann, R., Eswar, N., Penczek, P.A., Sali, A., Blobel, G., and Frank, J. (2001). Structure of the 80S ribosome from Saccharomyces cerevisiae—
tRNA-ribosome and subunit-subunit interactions. Cell 107, 373–386.
Triana-Alonso, F.J., Dabrowski, M., Wadzack, J., and Nierhaus, K.H. (1995). Self-coded 30-extension of run-off transcripts produces aberrant products during
in vitro transcription with T7 RNA polymerase. J. Biol. Chem. 270, 6298–6307.
Tung, C.S., Joseph, S., and Sanbonmatsu, K.Y. (2002). All-atom homology model of the Escherichia coli 30S ribosomal subunit. Nat. Struct. Biol. 9, 750–755.
Waterhouse, A.M., Procter, J.B., Martin, D.M., Clamp, M., and Barton, G.J. (2009). Jalview Version 2—a multiple sequence alignment editor and analysis work-
bench. Bioinformatics 25, 1189–1191.
Whitford, P.C., Ahmed, A., Yu, Y.A., Hennelly, S.P., Tama, F., Spahn, C.M.T., Onuchic, J.N., and Sanbonmatsu, K.Y. (2011). Excited states of ribosome
translocation revealed through integrative molecular modeling. Proc. Natl. Acad. Sci. USA 108, 18943–18948.
Cell 158, 121–131, July 3, 2014 ª2014 Elsevier Inc. S3
Figure S1. Heterogeneity in the 80S Ribosome POST Complexes from Rabbit Liver, Related to Figure 1
(A) Top: The POST 80S complex (initial data set is 1,072,632 images) was analyzed and sorted into distinct classes according to multi-particle alignment
procedures: POST (236,113 images), PRE-classical-1 (191,718 images), PRE-rotated-2 (100,473 images) and eEF2-containing (143,150 images) (middle).
Subsequent refinement of the POST and PRE (classical-1) classes yielded four distinct structures: POST (236,113 images), PRE (classical-1) (66,618 images),
PRE (classical-2) (73,951 images) and a structure with fragmented A-site tRNA (51,149 images) which is not in the scope of this study (bottom).
(B) Fourier Shell Correlation (FSC) curves of the presented complexes. The inset table shows the final resolutions according to the FSC 0.5 criterion for each
complex.
(C) Cross-resolution curves between each complex and its atomic model indicate resolutions in the same range as the FSC analysis, thus corroborating our
resolution estimation.
S4 Cell 158, 121–131, July 3, 2014 ª2014 Elsevier Inc.
Figure S2. Eukaryotic-Specific Contacts between P-tRNA and Large Ribosomal Proteins, Related to Figure 2(A) Contact of the P-site tRNA (position 1) with ribosomal protein uL16 (L10e). The same contact of uL16 (L10e) and the peptidyl-tRNA in the A/P-site has been also
observed in the rotated-2 subpopulation of the PRE ribosome (Budkevich et al., 2011). Thus, the uL16 loopmight escort the 50-end of peptidyl-tRNA from the A/P-
hybrid position to the classical P/P-site of the POST state.
(B) Contacts of the P-tRNA elbow with ribosomal proteins eL44 and uL5 (rpL11). The so-called P-site loop (Rhodin and Dinman, 2010), a highly conserved,
basically charged internal loop of uL5 (amino acids 48 to 68) appears to participate in this interaction and also in a nearby interaction with the D-loop of the P-site
tRNA.
White arrows indicate contacts. Themodels for tRNAs and ribosomal proteins are derived from the homology model of the human 80S ribosome presented in this
paper. Ribosome orientations are indicated by the orientation aids (top left).
Cell 158, 121–131, July 3, 2014 ª2014 Elsevier Inc. S5
Figure S3. 40S Rolling and Its Effect on Integrity of Ribosomal Intersubunit Bridges and on tRNA Positions, Related to Figure 3
(A) Comparison of the 80S ribosome in fully ratcheted state (Spahn et al., 2004b) with the ribosome in classical-1 state (this study). The fully ratcheted state is
characterized by a rotation of the 40S subunit of 5� and a swivel movement of the head of 14�.(B) Comparison of the ribosome in classical-1 state (this study) to the ribosome in rotated-2 state (Budkevich et al., 2011) shows a rotation of the 40S subunit of 8�.(C) Comparison of the 40S subunit positions in classical-1 state to the POST state (both from this study) reveals a new motion of 40S subunit of 6� we term
‘‘subunit rolling.’’ Comparison of the 40S positions in classical-1 and POST states is already shown in the related Figure 3 but color-coded from 0�20 A instead of
0 �12 A.
(D) Intersubunit bridges of the POST complex. In keeping with the previously established nomenclature (Spahn et al., 2001), universally conserved bridges are
numbered B1-B7 (red) and eukaryotic-specific are numbered eB8-eB11 (pink). Furthermore, additional bridges eB12-eB14 that were described for yeast 80S
ribosomes in (Ben-Shem et al., 2011) are also observed. Bridges B6, B7 and eB13 (green) were not observed in the POST state due to subunit rolling. Moreover,
bridge eB8 (eS1(S3A)/H79; green) is absent in the POST state as well because it is a property of the 80S ribosome in the rotated conformation (Ben-Shem et al.,
2011).
(E) Comparison of the positions of the eukaryotic PRE and POST tRNAs near the elbow regions as observed in mammalian system (colored ribbons) with those
observed in T. thermophilus 70S structures (Voorhees et al., 2010), (gray ribbon, PDB ID code 2WDG). Structures of P- and E-site tRNAs along with their paired
mRNA codons were docked independently as rigid bodies to the PRE and POST cryo-EM maps. Color code: P-tRNA in PRE complex (blue), P-tRNA in POST
complex (green), E-tRNA in PRE complex (red), E-tRNA in POST complex (orange). Distances were measured between phosphate backbones at position 56.
(F) Effects of subunit rolling on the tRNA positions. Structures as in (E) but color-coded by distances from 0 to 5 A.
S6 Cell 158, 121–131, July 3, 2014 ª2014 Elsevier Inc.
Figure S4. Protein Sequence Alignment of EF1A Homologs from Different Species, Related to Figure 4
(A) A protein sequence alignment of EF1A homologs from different species clearly demonstrates that, while EF1A/EF-Tu sequences are largely conserved,
archaeal and eukaryotic homologs feature a conserved helical extension of the switch I region (highlighted yellow in the structural annotation). This insertion is
located between the conserved a-helix 1 and b strand 2 and is not present in EF-Tu. The sequence alignment was calculated using Clustal omega (Sievers et al.,
2011) with sequences obtained from the UniProt database (Apweiler et al., 2004) (IDs Q9YAV0, P68104, P68103, P60338, P53013, P50522, P10126, P0CE47).
Different from the initial E. coli EF-Tu1 structure (PDB ID code 1ETU), we used the UniProt numbering including the leading methionine. This results in a +1
sequence shift, such that e.g.,E. coliH84 is H85 according to our numbering. Visualization of the sequence alignment according to theClustalX-color scheme and
a representation of their phylogenetic relationship were prepared using JalView (Waterhouse et al., 2009). For the consensus sequence, functional regions are
highlighted in color (green: P loop, red: switch I region, blue: switch II region) and the corresponding GTP-binding sequence motifs (Kjeldgaard et al., 1996) in
white.
(B) Location of the eukaryotic-specific insertion (helix 5) between domain I and II. The density corresponded to the helix 5 of rabbit eEF1A is highlighted in pink,
while the corresponding section of the A. pernix X-ray model (PDB ID 3VMF) is shown in dark blue.
S8 Cell 158, 121–131, July 3, 2014 ª2014 Elsevier Inc.
Figure S5. Reorganization of the Mammalian Ternary Complex to Adopt to a New Ribosomal Environment, Related to Figure 5
(A) Structural and sequence comparison of the interface between domain III of aEF1A/EF-Tu and the acceptor (stem and elbow of A/T-tRNA for archaeal (blue)
and bacterial (gold) X-ray models (bacterial ternary complex: PDB ID 2XQD; archaeal EF1A: PDB ID 3VMF; A/T-tRNA: hybrid structure from PDB IDs 2XQD for the
ASL and 1TTT for the tRNA body). Analysis of atom-atom distances between the bacterial tRNA and the archaeal EF1Amodel reveals multiple clashes (indicated
by red lines in the structure and red-orange bars in the sequence panel). Most likely, several insertions (dark blue) compared to the bacterial EF-Tu extend EF1A
either directly or indirectly (light blue) toward the bacterial tRNA position. As a consequence, the position of the archaeal tRNAmust differ from that of its bacterial
counterpart due to steric hindrance. Interactions of the aEF1A with the A/T-tRNA are formed by the loops around His335/Pro336 (archaeal numbering) and
Asp414/Met415 archaeal numbering) around positions tRNA 52 and 63/64, respectively. Orientation aids between the structure and the sequence are given by
small letters highlighting regions of interest (a, b, c and d). The structure-based sequence comparison was calculated using Chimera (Pettersen et al., 2004).
(B) Superposition of the GTP-binding domains of archaeal EF1A,GTP (Kobayashi et al., 2012; PDB ID code 3VMF; red ribbon) and yeast eEF1A (Andersen et al.,
2000; PDB ID code 1F60; gray ribbon). The helical extension (aminoacids 35-50) is depicted in orange. Conserved switch I helixes (aminoacids 57-61) and
(aminoacids 63-68) are colored yellow while switch II region is depicted blue.
Cell 158, 121–131, July 3, 2014 ª2014 Elsevier Inc. S9